|
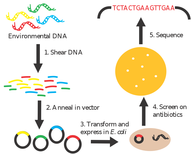
Recent studies of natural environments have revealed vast genetic reservoirs of antibiotic resistance (AR) genes. While antibiotic resistant bacteria have been cultured from seawater, we currently have little knowledge of the frequency and genetic diversity of AR genes in marine environments. To address this, I used functional metagenomics, a random shotgun cloning technique, to clone fragments of DNA from marine environments. I then screened these clones on several antibiotics to test for resistance. Antibiotic resistant clones were found in all locations but with varying frequency. Of the sequenced clones, 35% could be indentified as known AR genes including beta-lactamases and bicyclomycine resistance pumps. In addition, we isolated genes that conferred antibiotic resistance but could not be identified as previously classified AR genes. However, the functions were similar to known AR genes (e.g. transporters, hydrolases, oxioreductases). Furthermore, 44% of genes conferring antibiotic resistance were found in marine taxa (e.g., Pelagibacter sp. Polaribacter sp., Prochlorococcus, Roseobacter sp., Vibrio sp., etc.). This study demonstrates a previously unknown wide diversity and distribution of antibiotic resistance genes among marine environments. Thus ocean environments represent a global reservoir of both clinically relevant and potentially novel antibiotic resistance genes.
|
Where are species located has been a central question in biology. While the focus has mainly been on the distributions of plants and animals, distribution patterns of microorganisms is becoming well established. My current work answers the question of when are species located by investigating the temporal patterns of marine microbial diversity. To address this, I have been in charge of the Microbes of the Coastal Region of Orange County (MiCRO) time series at Newport Pier, Newport Beach, CA in collaboration with Steve Allison, Jennifer Martiny, and Sunny Jiang.
I am using high-throughput sequencing of the RNA polymerase (rpoC1) marker gene to identify the composition of cyanobacterial communities over a four-year time series. In addition, I have been using data from databases like the International Census of Marine Microbes and a collaboration with Jed Fuhrman at USC to look at the effect of temporal scale on bacterial beta-diversity. We found that beta-diversity does depend on temporal scale. While communities do turnover at seasonal scales, there is considerable turnover at intra-seasonal scales. Hatosy, S.M., J.B.H. Martiny, R. Sachdeva, J. Steele, J.A. Fuhrman, A.C. Martiny. 2013. Beta-diversity of marine bacteria depends on scale. Ecology. 94:1894-1904 Publications related to the MiCRO time series: Allison, S, Y. Chao, J.D. Fararra, S. Hatosy, A.C. Martiny. 2012. Fine-scale temporal variation in marine ectoenzymes of coastal southern California. Frontiers in Microbiology. doi:10.3389/fmicb.2012.00301 Freitas, S, S. Hatosy, J.A. Fuhrman, S.M. Huse, D.B.M. Welch, M.L. Sogin, A.C. Martiny. 2012. Global distribution and diversity of marine Verrucomicrobia. ISME J. 6:1499-1505 |